Powerful new molecular tools are making genetic engineering faster, easier, and more precise than ever before, raising new possibilities for researchers and clinicians who seek to study, or even alter, the course of disease. In a lecture last week at Harvard Medical School (HMS), J. Keith Joung ’87, M.D.-Ph.D. ’96, associate chief of pathology for research at Massachusetts General Hospital and associate professor at HMS, summarized recent developments in the rapidly developing field of precision genome editing, which the journal Science named a runner-up for its 2012 Breakthrough of the Year (the winner was the discovery of the Higgs boson). The new technologies, which Science dubbed “genomic cruise missiles,” enable researchers to insert or knock out genes by targeting gene-altering proteins to specific DNA sequences.
Recent advances in molecular biology have given researchers an increasingly sophisticated ability to manipulate the genomes of large, multicellular organisms like plants and mice, often by adapting existing biological features. For instance, RNA interference, recognized by the 2006 Nobel Prize in physiology or medicine, allows biologists to temporarily disable specific genes by disrupting the translation of DNA instructions into proteins that carry them out. A year later, the Nobel committee recognized another set of genome-engineering techniques that modify embryonic stem cells to create “knockout” mice, in which particular genes have been altered or disabled.
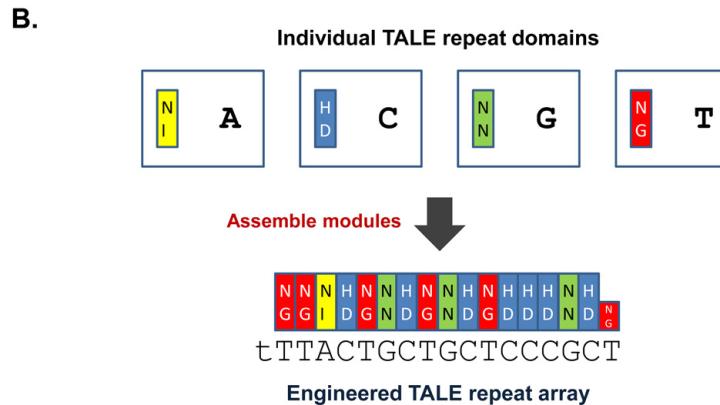
The new technologies, collectively referred to as “targeted nucleases” (a nuclease cuts a nucleic acid like DNA), promise to be more permanent than RNA interference, but less laborious than the methods required to create knockout mice. One technique makes use of bacterial proteins called transcription activator-like effectors (TALEs), which recognize and bind to DNA using a string of protein domains, each of which recognizes one of DNA’s four nucleotides—adenine, thymine, guanine, and cytosine. By linking several of these interchangeable DNA-binding domains in a row, along with a nuclease domain, the protein can be engineered to recognize and cut highly specific DNA sequences. After the DNA is cut, researchers can take advantage of DNA-repair mechanisms to make DNA modifications or introduce new genes.
The modularity of the DNA-binding domains makes TALE nucleases especially easy to design and manufacture. Joung’s lab, for instance, published a method in 2012 to engineer multiple customized proteins in just two weeks. Two of his recent papers, published this month in Nature Biotechnology, suggest that by swapping the nuclease for other protein domains—for instance, domains that modify biochemical markers that direct gene expression—the TALE proteins can be adapted for other purposes.
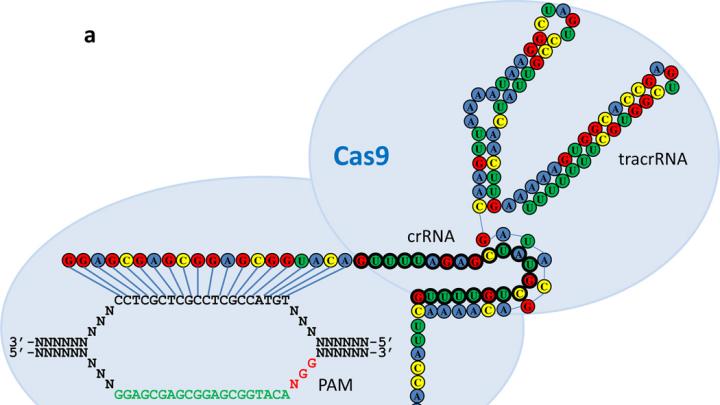
Another developing technology harnesses proteins involved in bacterial immune systems. When viruses attack, bacteria copy and store pieces of the attacking viral DNA in their own bacterial genomes. Generations later, if the viruses attack again, the bacterial DNA database recognizes the invading viral DNA and activates systems to destroy it. Recently, scientists have begun adapting the system’s targeted attack capacities for precision genome editing. An advantage of this method, named CRISPR-Cas after the proteins and DNA involved, is that it uses RNA to seek out target DNA sequences, rather than the DNA-binding protein domains found in TALEs; RNA is easier to manufacture than proteins.
The past year and a half has seen a flurry of advances in the development of CRISPR-Cas as researchers explore and refine its capabilities; Science titled an August news article “The CRISPR Craze.” In the Boston area, Winthrop professor of genetics George Church was among the first to successfully apply CRISPR-Cas genome editing to human cells, and Feng Zhang ’04, a core member of the Broad Institute and faculty member at MIT, has been able to modify multiple genes simultaneously. Joung’s lab showed that CRISPR, like TALEs, could be adapted to control gene expression.
Questions remain for the new technologies—Joung’s lab, in particular, has published studies suggesting that CRISPR-Cas may sometimes modify gene sequences outside of its intended target—but companies are already beginning to develop applications of the new technologies. Some researchers are reevaluating possibilities for gene therapy, which adds or modifies genes to treat disease. Early methods used viruses to insert therapeutic DNA into the genome, but the random delivery often had disruptive side effects. Targeted nucleases, in contrast, may offer a more precise delivery system. TALE nucleases and CRISPR-Cas methods are still under development, but several clinical trials are under way to test whether other targeted nucleases can treat HIV/AIDS by disabling the protein that HIV uses to enter human immune cells. The potential and limitations of targeted nucleases are still under investigation, but it seems clear that precision genome editing is here to stay.